Martijho-PathNet-search-plots
From Robin
(Difference between revisions)
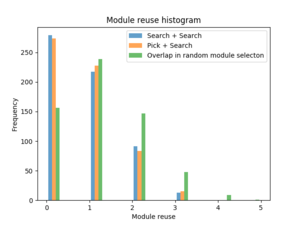
(Ny side: {|style="margin: 0 auto;" | [[File:600binMNIST_module_reuse_histogram.png|300px|thumb|upright|alt=module reuse plot|Distribution of module reuse in s+s and p+s searches along side the distr…) |
|||
| Line 1: | Line 1: | ||
| + | |||
| + | == Population Diversity == | ||
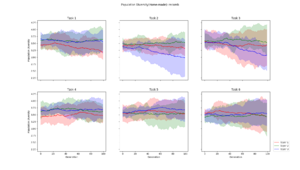
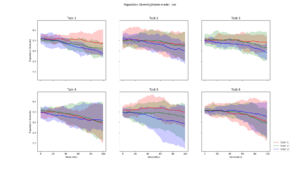
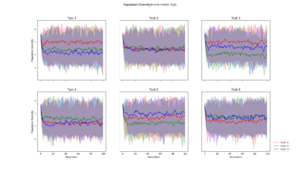
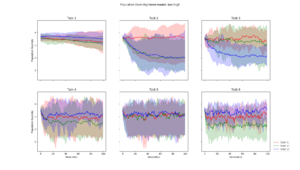
| + | Population diversity is calculated by the home-made metric of average distance to centroid genotype within each generation, which yields the same results as a pairwise Hamming distance | ||
| + | {|style="margin: 0 auto;" | ||
| + | | [[File:Population_diversity_recomb.png|300px|thumb|upright|alt=Population diversity in recomb search|FILLER]] | ||
| + | | [[File:Population_diversity_low.png|thumb|300px|upright|alt=Population diversity in low search|FILLER]] | ||
| + | | [[File:Population_diversity_high.png|thumb|300px|alt=Population diversity in high search|FILLER]] | ||
| + | | [[File:Population_diversity_low2high.png|thumb|300px|alt=Population diversity in low2high search|FILLER]] | ||
| + | | [[File:Population_diversity_high2low.png|thumb|300px|alt=Population diversity in high2low search|FILLER]] | ||
| + | |} | ||
| + | |||
{|style="margin: 0 auto;" | {|style="margin: 0 auto;" | ||
| [[File:600binMNIST_module_reuse_histogram.png|300px|thumb|upright|alt=module reuse plot|Distribution of module reuse in s+s and p+s searches along side the distribution for a random module selection]] | | [[File:600binMNIST_module_reuse_histogram.png|300px|thumb|upright|alt=module reuse plot|Distribution of module reuse in s+s and p+s searches along side the distribution for a random module selection]] | ||
Revision as of 15:35, 27 February 2018
Population Diversity
Population diversity is calculated by the home-made metric of average distance to centroid genotype within each generation, which yields the same results as a pairwise Hamming distance